June 29, 2017
 There’s a biological revolution underway and its name is CRISPR.
There’s a biological revolution underway and its name is CRISPR.
Pronounced ‘crisper’, the technique stands for Clustered Regularly Interspaced Short Palindromic Repeat and refers to the way short, repeated DNA sequences in the genomes of bacteria and other microorganisms are organised.
Inspired by how these organisms defend themselves from viral attacks by stealing strips of an invading virus’ DNA, the technique splices in an enzyme called Cas creating newly-formed sequences known as CRISPR. In bacteria, this causes RNA to make copies of these sequences, which help recognise virus DNA and prevent future invasions.This technique was transformed into a gene-editing tool in 2012 and was named Science magazine’s 2015 Breakthrough of the Year. While it’s not the first DNA-editing tool, it has piqued the interest of many scientists, research and health groups because of its accuracy, relative affordability and far-reaching uses. The latest? Eradicating HIV.
At the start of May, researchers at the Lewis Katz School of Medicine at Temple University (LKSOM) and the University of Pittsburgh demonstrated how they can remove HIV DNA from genomes of living animals – in this case, mice – to curb the spread of infection. The breakthrough was the first time replication of HIV-1 had been eliminated from infected cells using CRISPR following a 2016 proof-of-concept study.
In particular, the team genetically inactivated HIV-1 in transgenic mice, reducing the RNA expression of viral genes by roughly 60 to 95 per cent, before trialling the method on infected mice.
“During acute infection, HIV actively replicates,” Dr. Khalili explained. “With EcoHIV mice, we were able to investigate the ability of the CRISPR/Cas9 strategy to block viral replication and potentially prevent systemic infection.”
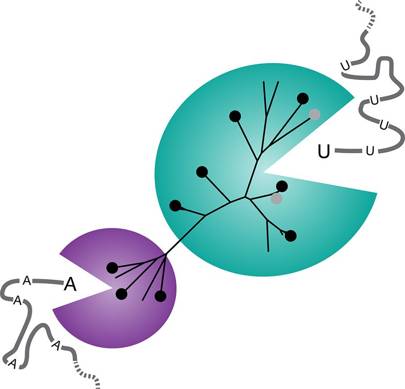
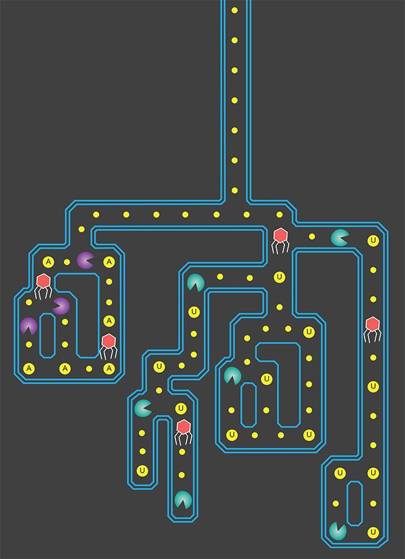
Since the HIV research was published, a team of biologists at University of California, Berkeley, described 10 new CRISPR enzymes that, once activated, are said to “behave like Pac-Man” to chew through RNA in a way that could be used as sensitive detectors of infectious viruses.
These new enzymes are variants of a CRISPR protein, Cas13a, which the UC Berkeley researchers reported last September in Nature, and could be used to detect specific sequences of RNA, such as from a virus. The team showed that once CRISPR-Cas13a binds to its target RNA, it begins to indiscriminately cut up all RNA making it “glow” to allow signal detection.
Two teams of researchers at the Broad Institute subsequently paired CRISPR-Cas13a with the process of RNA amplification to reveal that the system, dubbed Sherlock, could detect viral RNA at extremely low concentrations, such as the presence of dengue and Zika viral RNA, for example. Such a system could be used to detect any type of RNA, including RNA distinctive of cancer cells.
This piece has been updated to remove copy taken from WIRED US.